Example: psbA-trnH is the genomic region of interest
1. Go to http://plantaligdb.portugene.com/cgi-bin/PlantAligDB_home.cgi
In the main page, you can access the section Genomic Regions on the bar
2. Click on the button Genomic Regions
In this section you find the genomic regions present in PlantAligDB
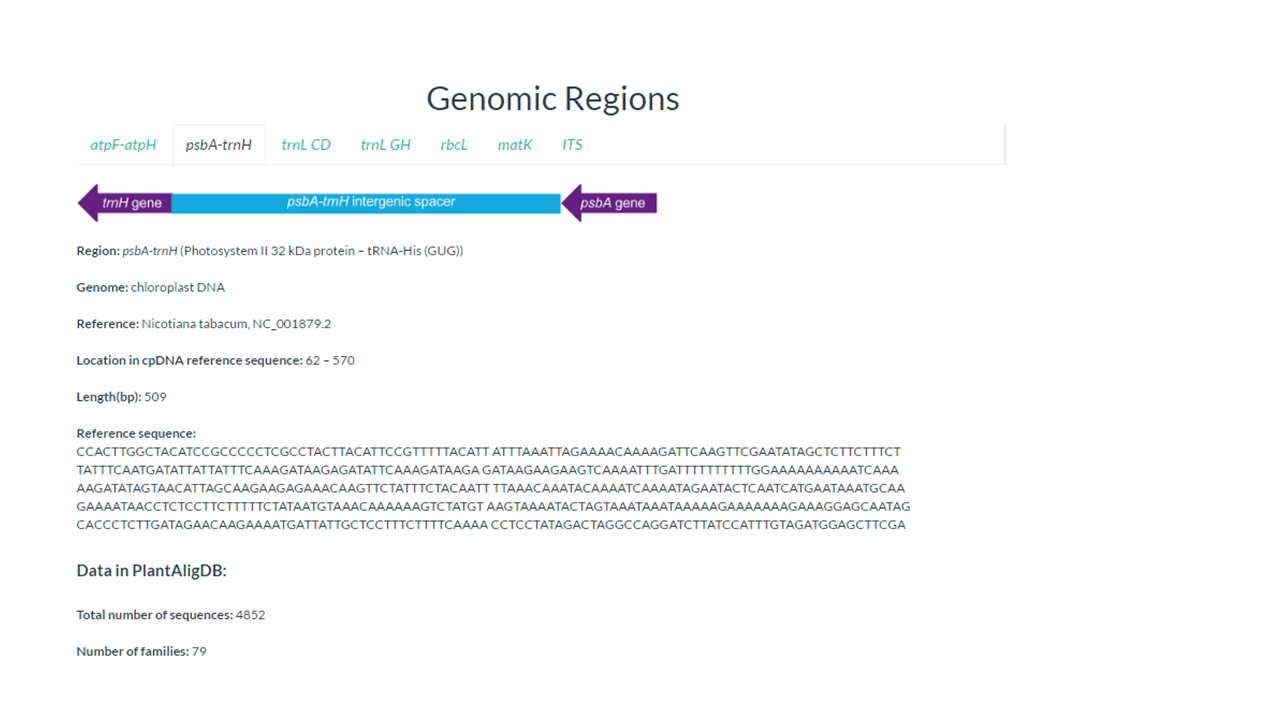
3. Click on psbA-trnH
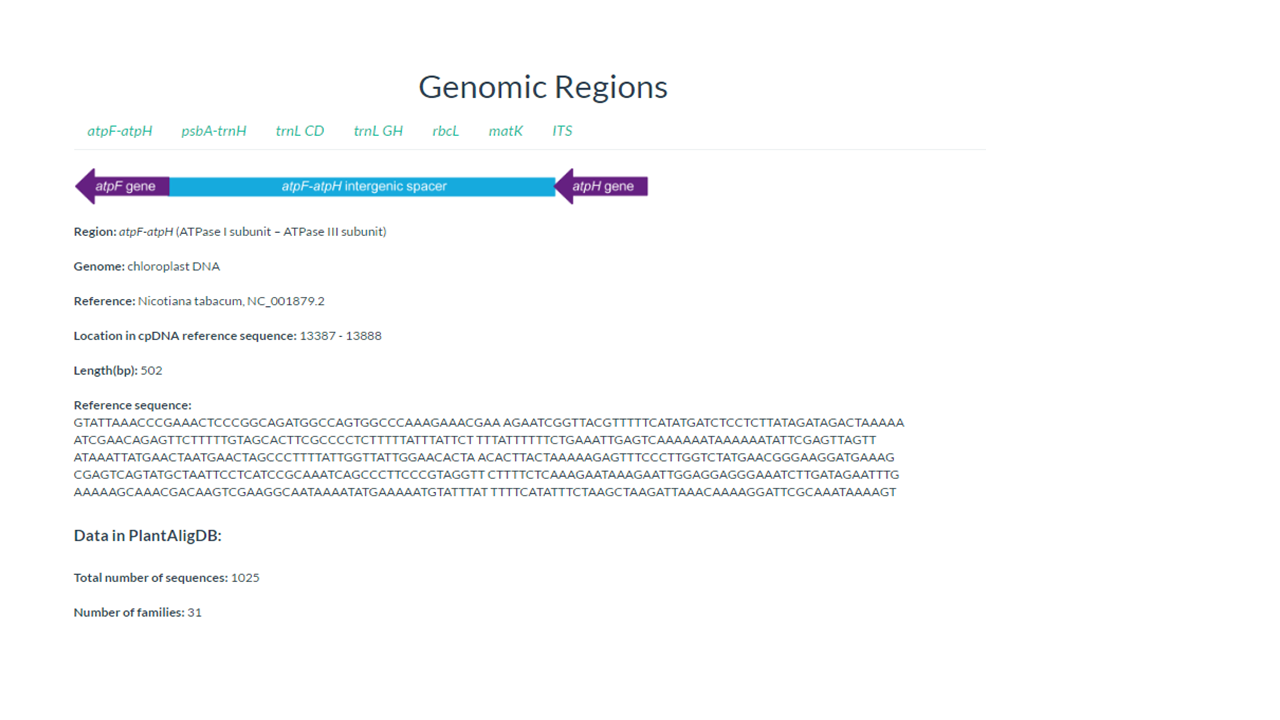
In this page you find information about the target genomic region
In our example psbA-trnH
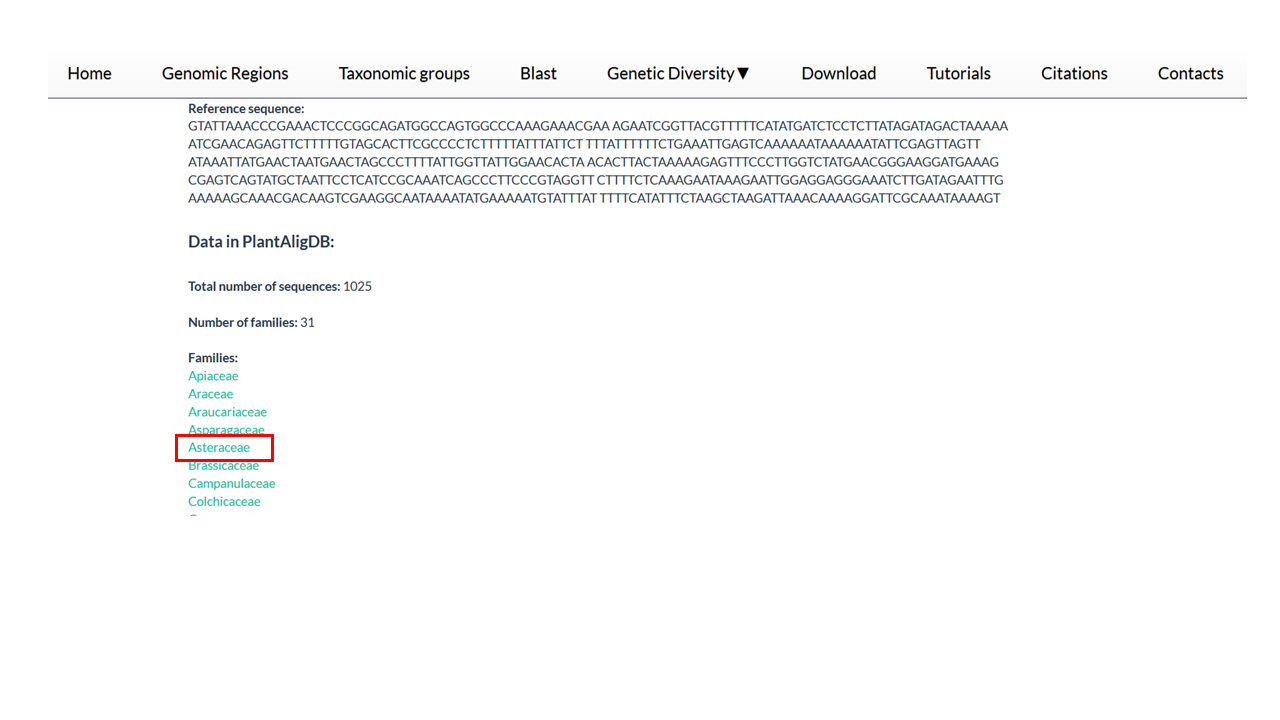
Below the description of the genomic region you find the families organized in alphabetical order.
4. Click on the name of a interested family
In our example, Asteraceae.
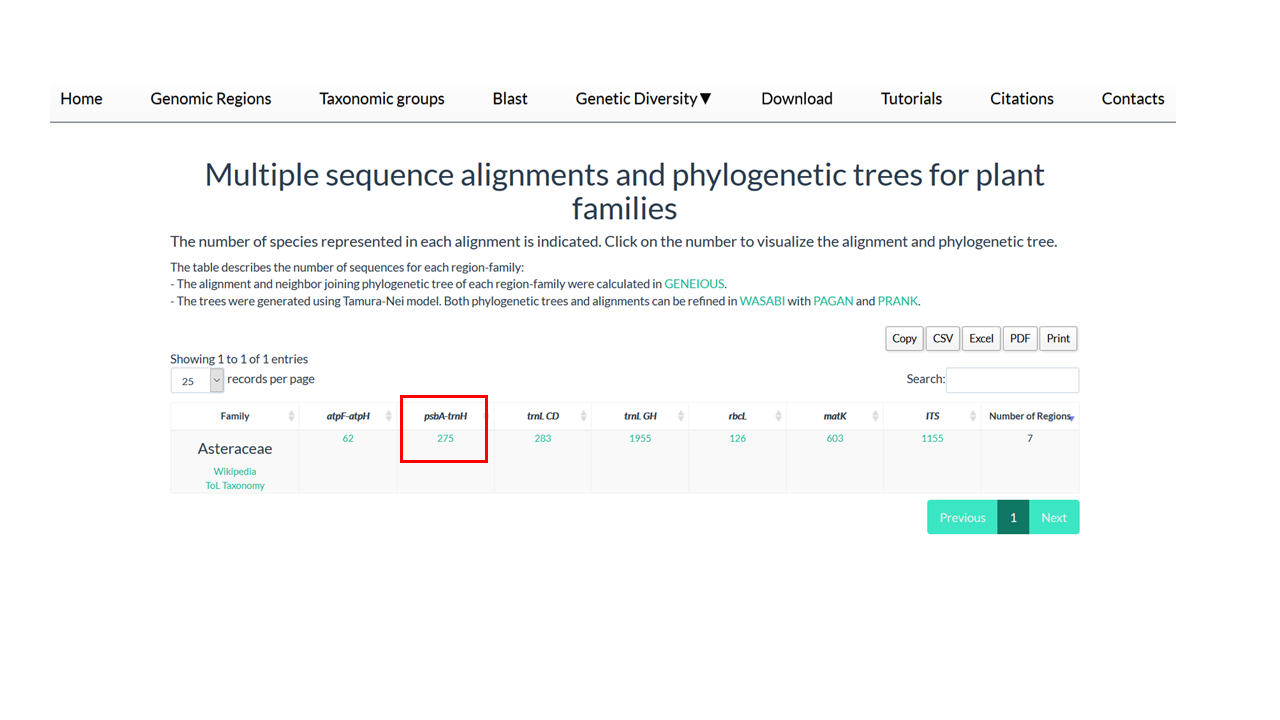
5. Clik on the number below of target region.
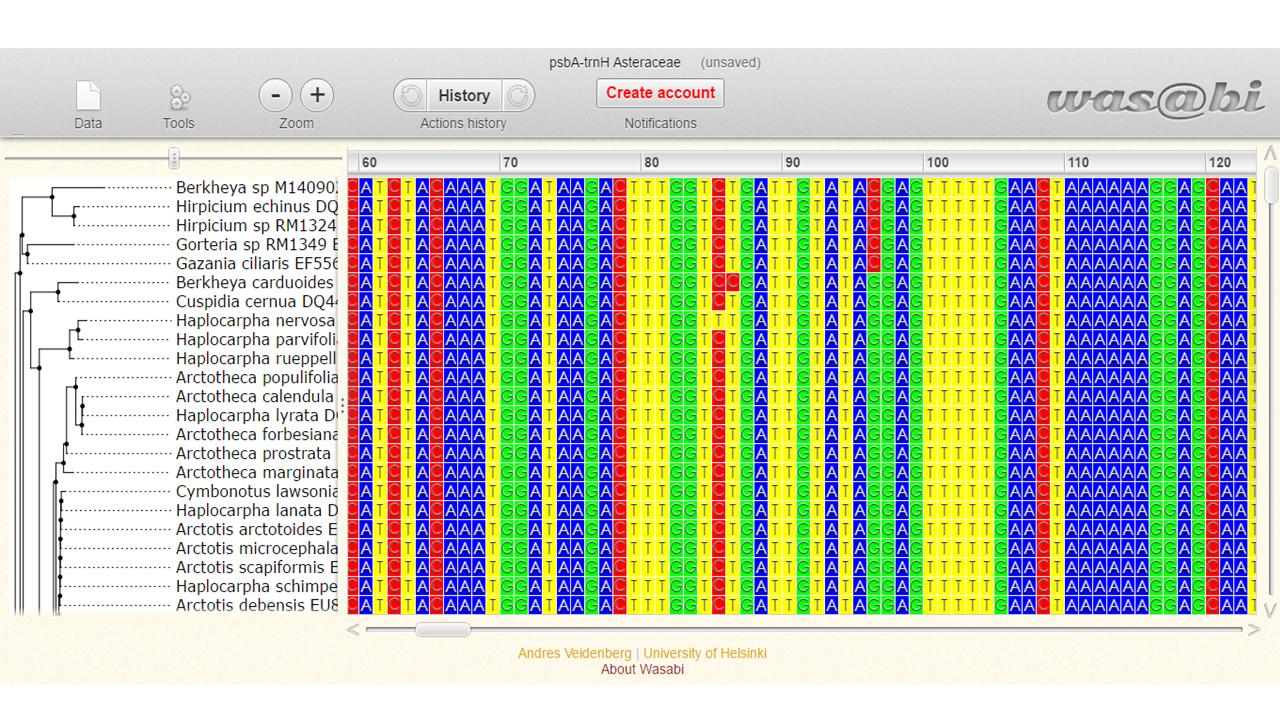
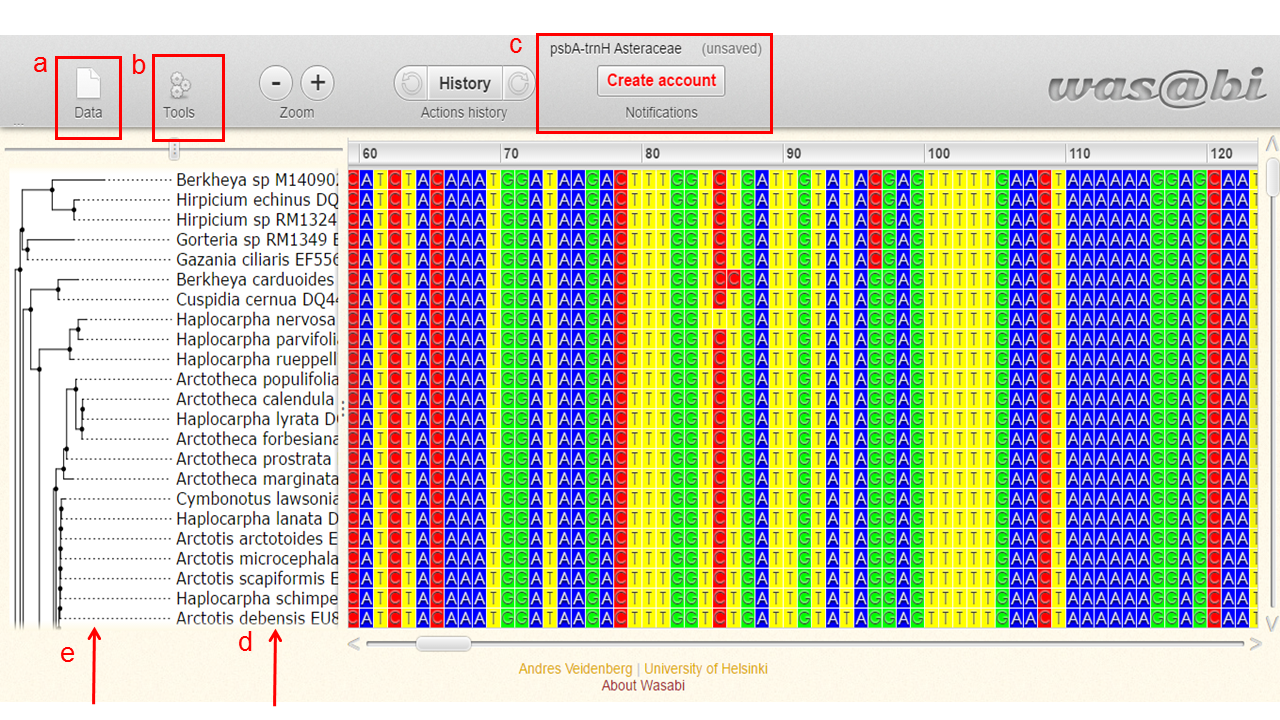
You will be direct for a visualization tool called Wasabi.
Here we can:
a. see information about the data and export them in fasta format
b. hide columns with gaps and edit some preferences
c. rename, save and export data
d. view the acession number and name of the species present in the alignment
Example: Solanaceae family
In the main page, you can access the section Taxonomic Groups on the bar
1. Click on the button Taxonomic Groups
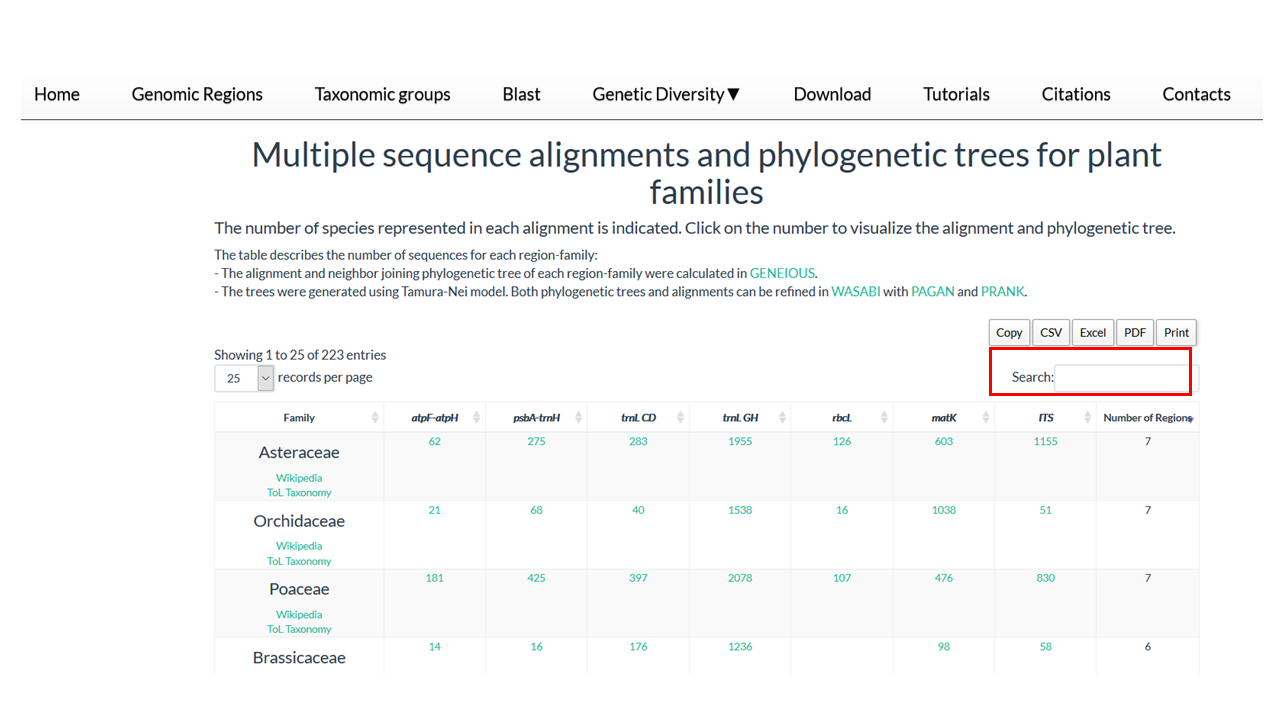
In this section you find the Taxomic Groups present in PlantAligDB
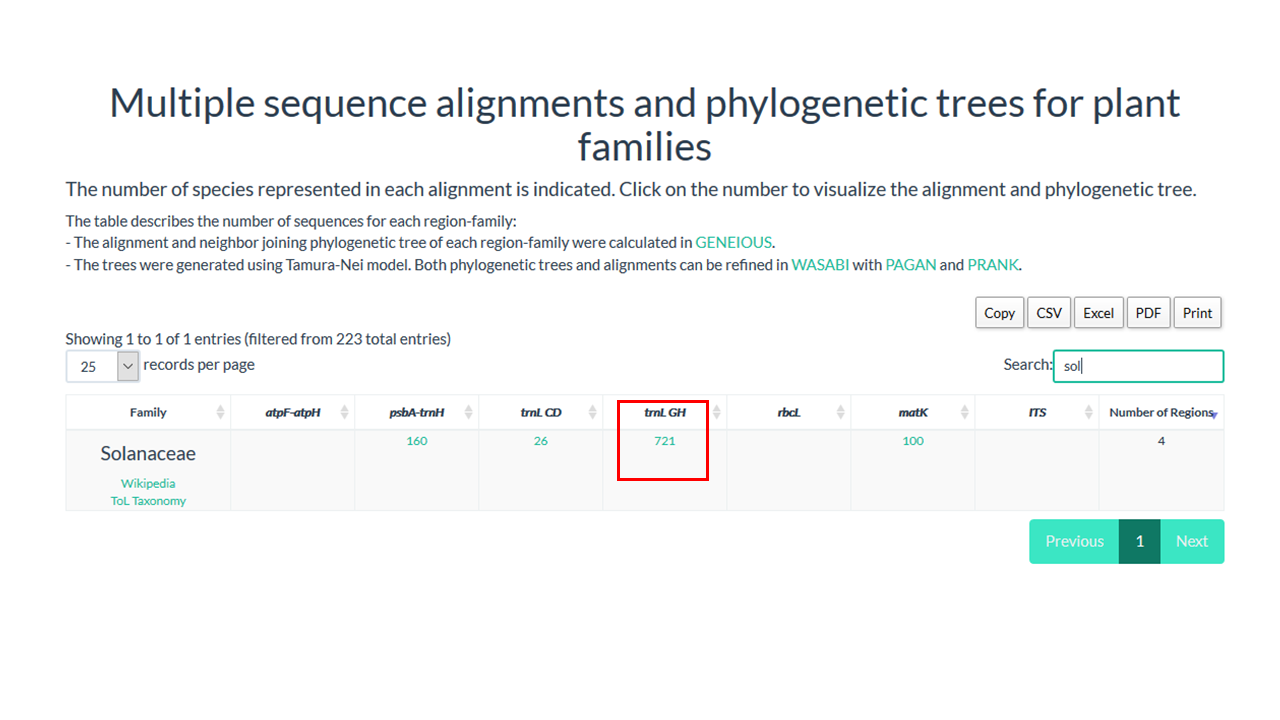
2. Go to the box 'Search' on right site
3. Type the name of the family
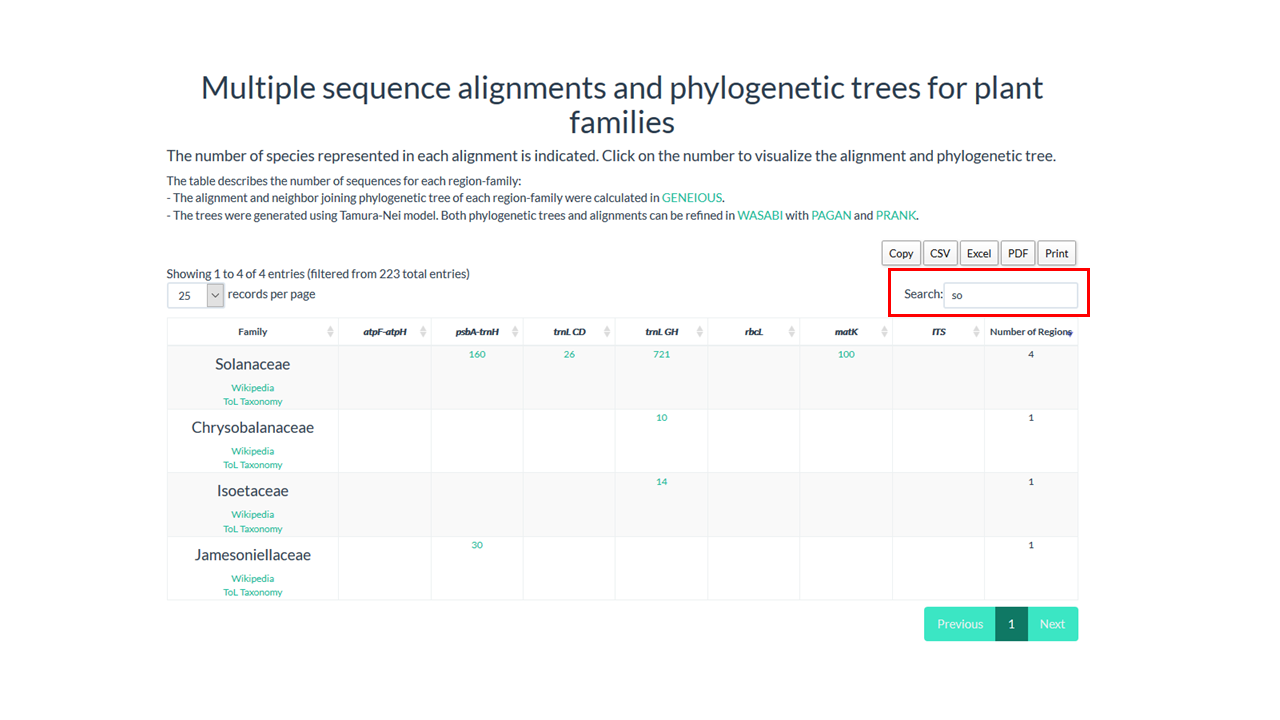
The dinamic table show if the family is present in database and the number of sequences in the alignment
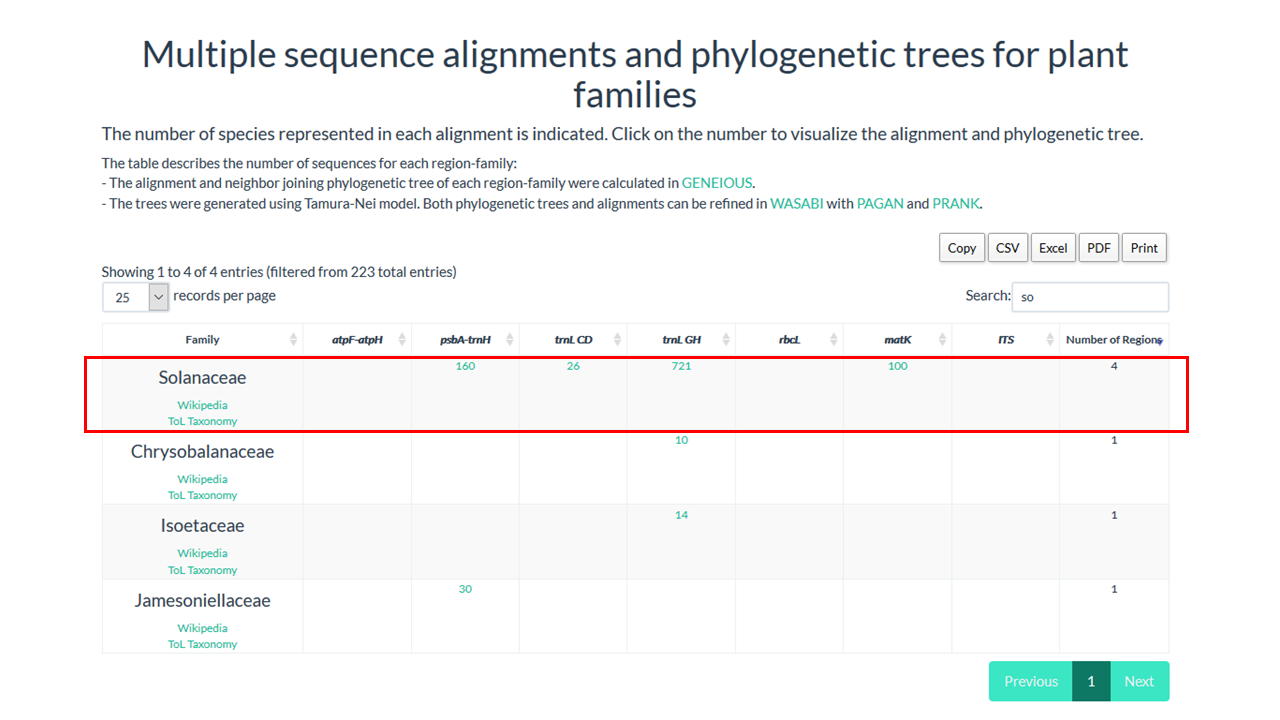
4. Click on the number below the region that you want to see the alignment
In our example Solanaceae have 721 sequences in the alignment for trnL GH region
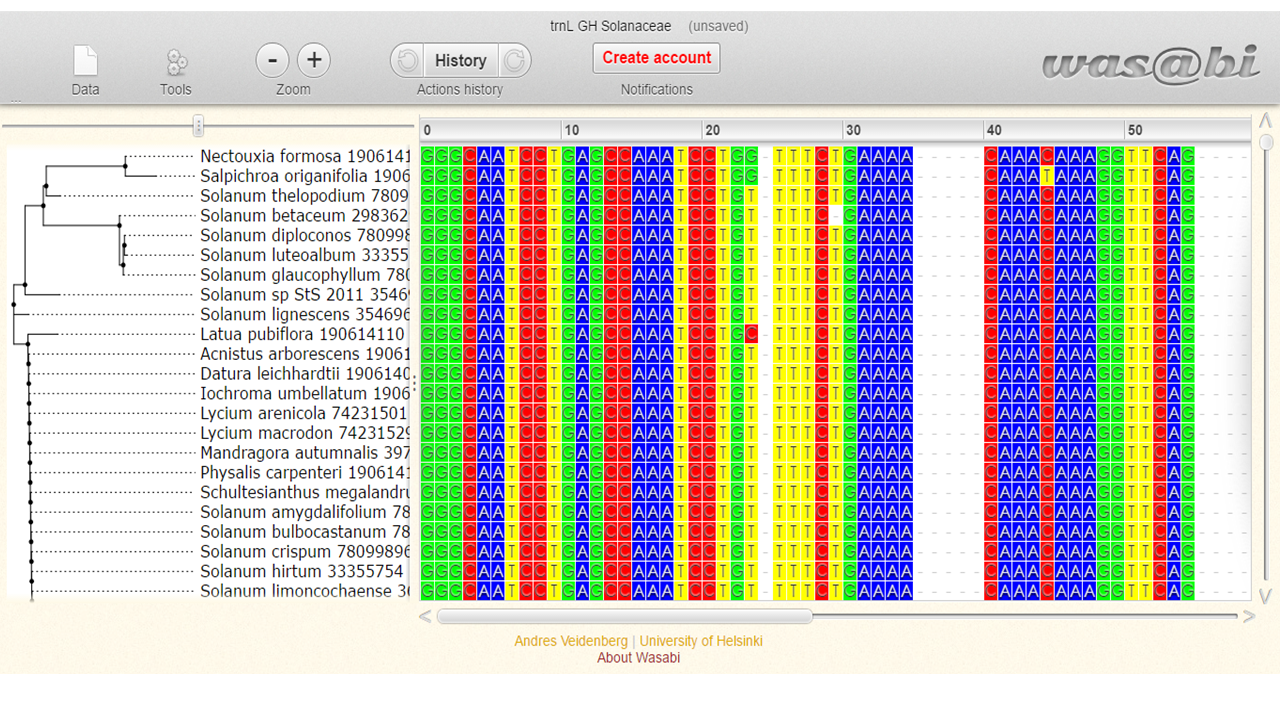
The alignment will be shown in Wasabi tool
In the main page, you can access the section BLAST on the bar
1. Click on the button BLAST
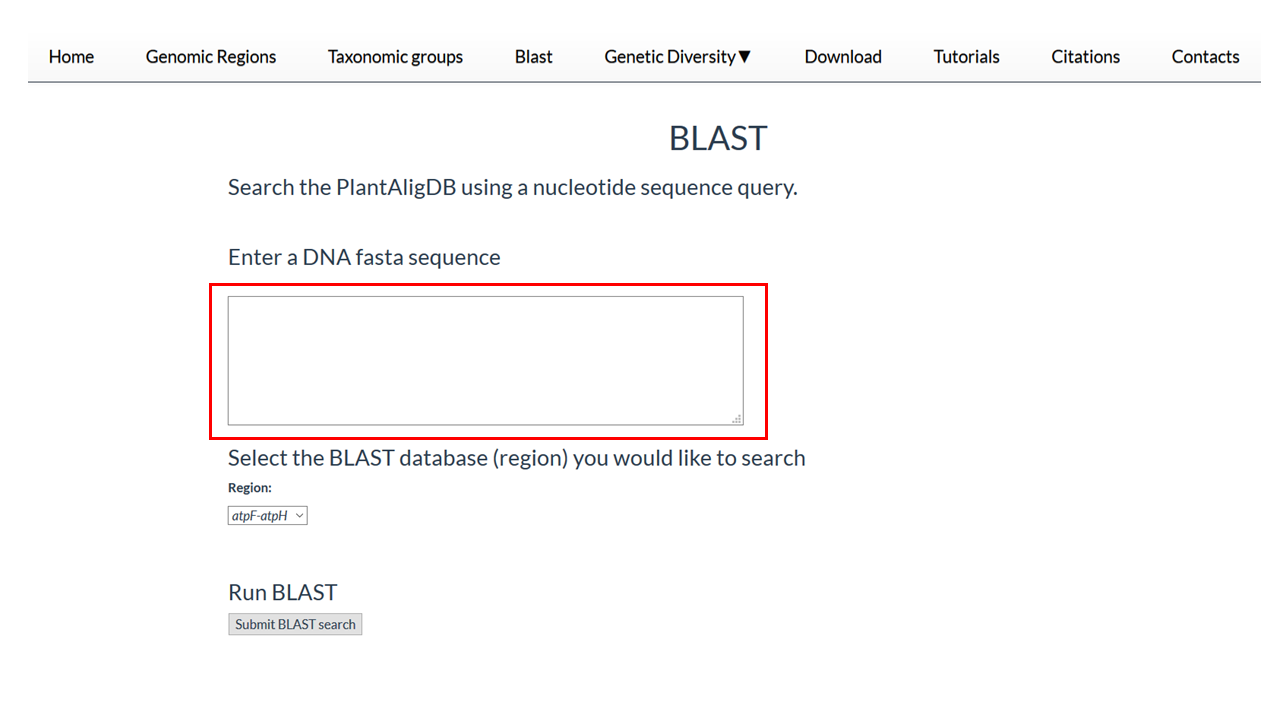
In this section you can verify the presence of a query sequence in PlantAligDB
2. Copy your DNA fasta sequence for the box
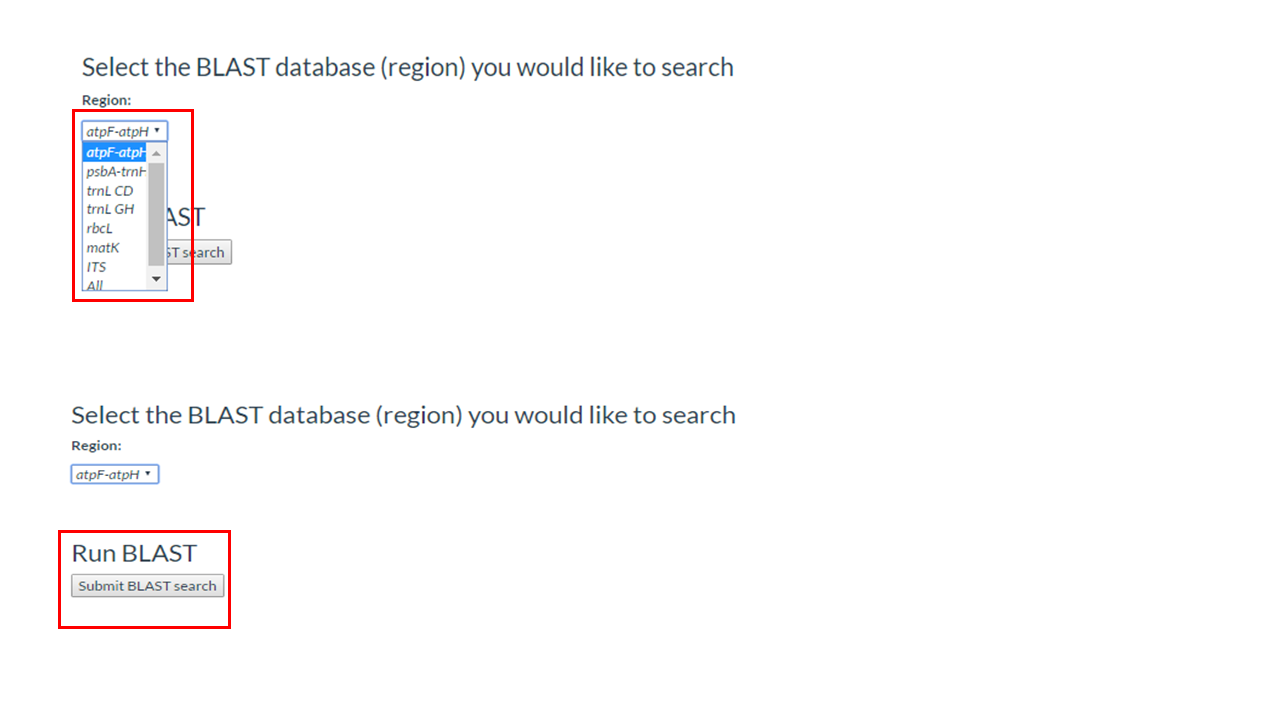
3. Select the BLAST database you would like to search or chose the option search against all nucleotide sequences (All)
4. Click on the button Submit BLAST search
You can generate automatic alignments from regions/families present in the database by using the following script:
--> Download
The script is used by us to update automatically the database.